Exploring the protein zoo
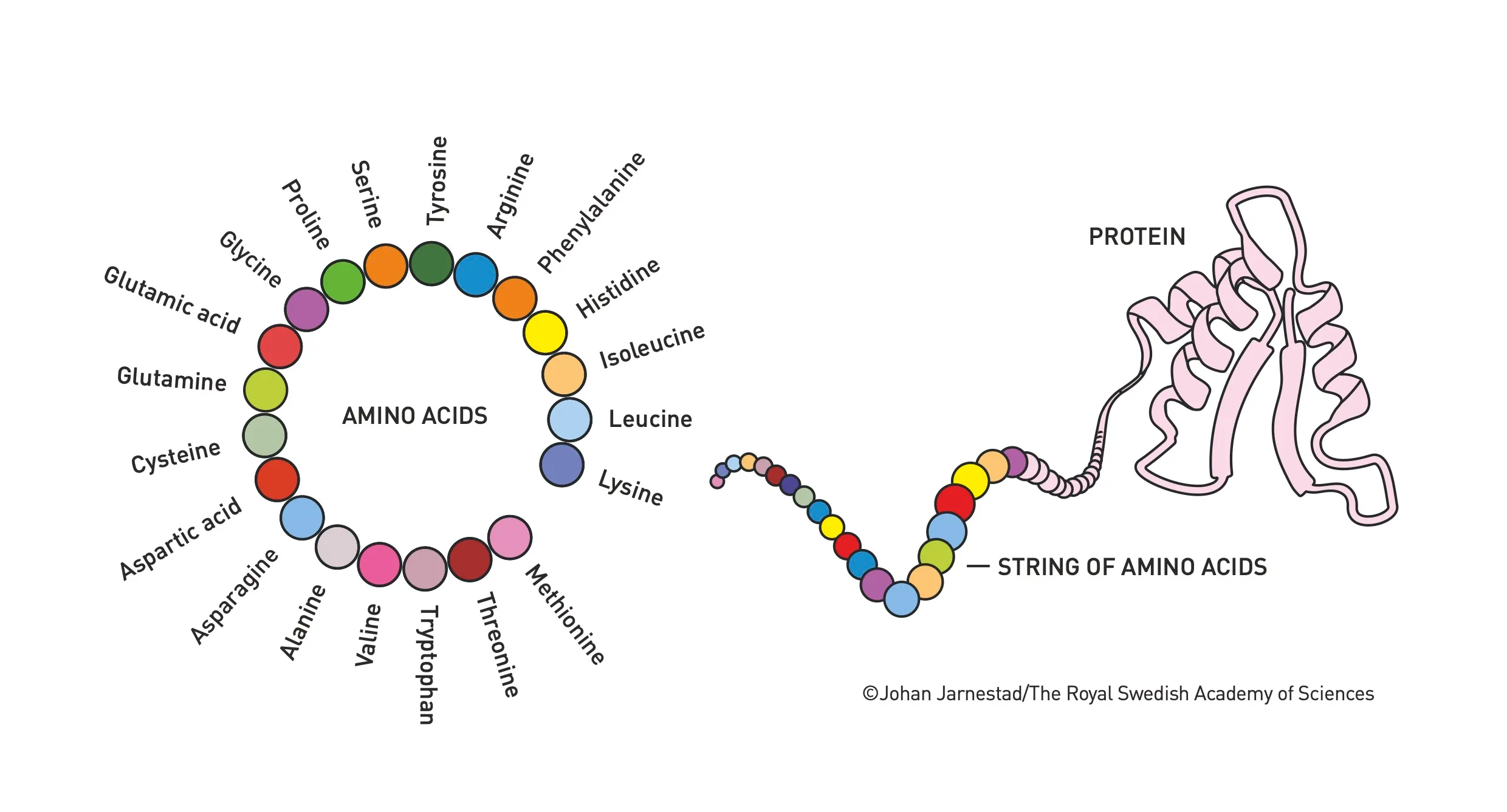
A protein is a chain of amino acids; its sequence largely determines its three-dimensional structure and biological function. From a library of just 20 standard amino acids, evolution over billions of years has produced an enormous diversity of proteins (the UniProt Knowledgebase contains more than 200 million entries!). Relating sequence to structure has been a grand challenge in biology for many decades, and efforts to address this problem were recognized with the 2024 Nobel Prize in Chemistry.
Below, you can load and explore both natural and designed protein structures (and other biological macromolecules – see the DNA origami examples) in the browser using 3Dmol.js. The viewer in the first section pulls coordinates from the RCSB Protein Data Bank by PDB ID. You can check out the PDB “Molecular Machinery” tour to find some other interesting examples. In the second section, you can either pull coordinates from the AlphaFold Database (given a UniProt ID) or load a structure directly from PDB text derived from a separate folding run. You can also export the structure as a .pdb file, which you can load in your external viewer of choice (e.g. Avogadro).
Structures from the RCSB Protein Data Bank
Enter a four-letter PDB ID or choose from one of the examples, then Load. To find other PDB IDs, you can search by protein name on the PDB website. Use the Style menu to switch rendering style.
Computationally predicted structures
There are two ways to get a model into the viewer below.
-
AlphaFold Database. If the AlphaFold Database already has a structure for your protein, enter a UniProt ID (e.g.
P42212for green fluorescent protein) and click Load. -
Paste PDB text. Run a computational sequence-to-structure pipeline elsewhere (e.g. using this ColabFold AlphaFold2 notebook), then paste the contents of the exported
.pdbinto the box and load.
Useful references
- H. M. Berman et al., The Protein Data Bank. Nucleic Acids Research (2000).
- J. Jumper et al., Highly accurate protein structure prediction with AlphaFold. Nature (2021).
- N. Rego & D. Koes, 3Dmol.js: molecular visualization with WebGL. Bioinformatics (2015).